Publications
Research articles
- Chen, Y. et al. Fica, S. M., et al., Whiffin, Nicola. (2024) De novo variants in the non-coding spliceosomal snRNA gene RNU4-2 are a frequent cause of syndromic neurodevelopmental disorders. Nature, 6322, 832-839.
Genomics identified the first snRNA variants directly linked to a neurological syndrome.
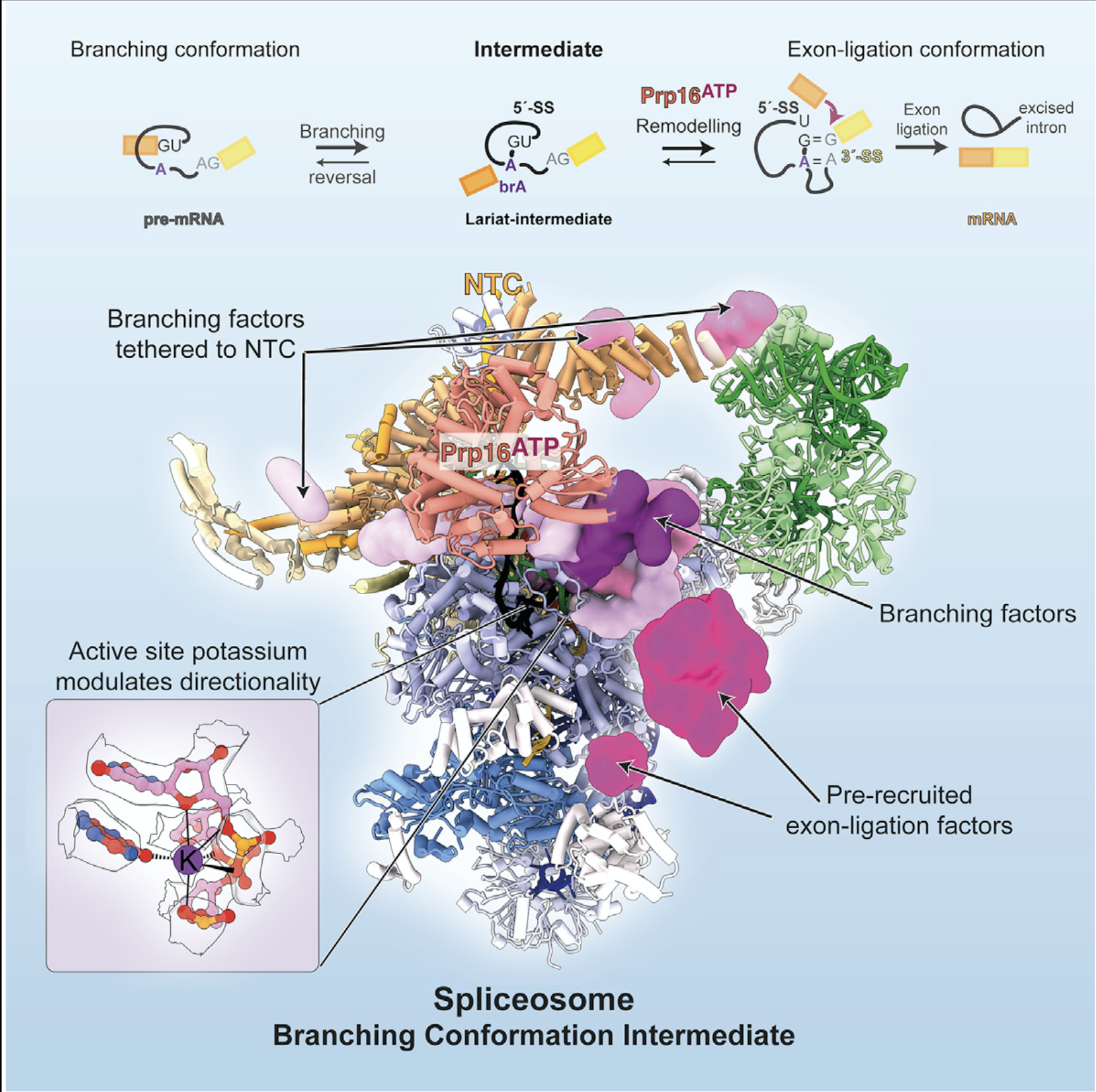
- Aupič, J., Borišek, J., Fica, S. M., Galej, W. P. & Magistrato, A. Monovalent metal ion binding promotes the first transesterification reaction in the spliceosome. Nat. Commun. 14, 8482 (2023).
Molecular dynamics identified a role for a specific K+ ion we discovered in the spliceosome active site.
- Parker, M. T., Fica, S. M., Barton, G. J. & Simpson, G. G. Inter-species association mapping links splice site evolution to METTL16 and SNRNP27K. eLife 12, e91997 (2023).
Phylogenetic analysis demonstrated how co-evolution of splicing factors and snRNA modifications enables plasticity of splice site selection during alternative splicing.
- Parker, M. T., et. al., Barton, G. J., Fica, S. M., Davies, B.H., Simpson, G. G. m6A modification of U6 snRNA modulates usage of two major classes of pre-mRNA 5’ splice site. eLife 11, e78808 (2022).
This study provided the first causal link between snRNA modifications and alternative splice site selection in plants.
- Strittmatter, L. M. Capitanchick, C., Halleger, M., Norman, C. M., Newman, A. J., Fica S. M.#, Oubridge, C., Luscombe, N.M. #, Ule, J. #, Nagai K. psiCLIP reveals dynamic RNA binding by DEAH-box helicases before and after exon ligation. Nat Commun 12, 1488 (2021). # corresponding authors
A new technique to study interactions of ATPases and other RNA-binding proteins with the substrate pre-mRNA in the fully assembled spliceosome uncovered a previously unappreciated role of spliceosomal ATPases in proofreading the stability of the spliceosome during mRNA synthesis.
- Wilkinson, M. E.*#, Fica S. M.*#, Galej, W. P.*, Nagai, K. (2021) Structural basis for conformational equilibrium of the catalytic spliceosome. Molecular Cell, 81, 1-14, *co-first authors, # corresponding authors.
Cryo-EM revealed that the spliceosome conformations for the two catalytic steps are in equilibrium during the catalytic stage, explaining reversibility of the splicing reaction and how specific factors could modulate subtle changes in splicing efficiency based on splice site sequences.
- Fica, S. M.#, Oubridge, C., Wilkinson, M. E., Newman, A. J., Nagai, K. # (2019) A human postcatalytic spliceosome structure reveals essential roles of metazoan factors for exon ligation. Science, 363, 710-714, #corresponding authors.
The first structure of a human spliceosome right after mRNA synthesis revealed how mammalian spliceosomes co-opted novel protein factors to stabilize 3'-splice site docking in the active site.
- Wilkinson, M. E., Fica S. M., Galej, W. P., Norman, C. M., Newman, A. J., Nagai, K. (2017) Postcatalytic spliceosome structure reveals mechanism of 3'-splice site selection. Science, 358, 1283–1288.
This was the first structure of a yeast post-catalytic spliceosome and revealed the mechanism for 3'-splice site docking at the active site.
- Fica, S. M. #, Oubridge, C., Galej, W. P., Wilkinson, M. E., Bai, X.-C., Newman, A. J., & Nagai, K. # (2017). Structure of a spliceosome remodelled for exon ligation. Nature, 542, 377-380, #corresponding authors.
This was the first structure of a yeast spliceosome in a conformation competent to perform ligation of the exons into mRNA and revealed how specific ATPases and exon-ligation factors promote the exon ligation conformation.
- Galej, W. P., Wilkinson, M. E., Fica, S. M., Oubridge, C., Newman, A. J., & Nagai, K. (2016). Cryo-EM structure of the spliceosome immediately after branching. Nature, 537, 197–201.
The first structure of a yeast spliceosome during the first step of catalysis revealed the structure of the active site, the mechanisms for branch point recognition, and the role of splicing factors in stabilising docking of the pre-mRNA in the active site.
- *Fica, S. M., *Mefford, M. A., Piccirilli, J. A., & Staley, J. P. (2014). Evidence for a group II intron-like catalytic triplex in the spliceosome. Nature Structural & Molecular Biology, 21, 464–471, *co-first authors.
This study demonstrated that the active site of the spliceosome is formed of an RNA triple helix.
- *Fica, S. M., *Tuttle, N., Novak, T., Li, N.-S., Lu, J., Koodathingal, P., Dai, Q., Staley, J. P., Piccirilli, J. A. (2013). RNA catalyses nuclear pre-mRNA splicing. Nature, 503, 229–234. *co-first authors.
This study demonstrated that the spliceosome active site is composed of RNA and not of protein.
- Schellenberg, M. J., Wu, T., Ritchie, D. B., Fica, Sebastian M., Staley, J. P., Atta, K. A., LaPointe, P. L., MacMillan, A. M. (2013). A conformational switch in PRP8 mediates metal ion coordination that promotes pre-mRNA exon ligation. Nature Structural & Molecular Biology, 20, 728–734.
This study demonstrated that an RNaseH-like motif in Prp8 is important for ligation of exons into mRNA.
Review articles
-
Pokorná, P., Aupič, J., Fica, S. M. & Magistrato, A. Decoding Spliceosome Dynamics through Computation and Experiment. Chem. Rev. (2025) doi:10.1021/acs.chemrev.5c00374.
- Parker, M. T., Fica, S. M. & Simpson, G. G. RNA splicing: a split consensus reveals two major 5′ splice site classes. Open Biol. 15, 240293 (2025).
A review of molecular mechanisms that modulate recognition and selection of 5’ splice sites during early stages of spliceosome assembly.
- Fica, S. M.# (2020). Cryo-EM snapshots of the human spliceosome reveal structural adaptions for splicing regulation. Current Opinion in Structural Biology, 65, 139–148. #corresponding author.
I reviewed the various structural snapshots of the human spliceosome and proposed that recruitment and substrate-specific engagement of human factors at different stages of the splicing cycle underlies increased plasticity of splice site selection by spliceosomes of complex organisms.
- Fica, S. M.#, & Nagai, K. # (2017). Cryo-electron microscopy snapshots of the spliceosome: structural insights into a dynamic ribonucleoprotein machine. Nature Structural & Molecular Biology, 24, 791–799, #corresponding authors.
An overview of recent spliceosome structures obtained by electron microscopy placed their findings in the context of previous genetic and biochemical studies.
- Nguyen, T. H. D., Galej, W. P., Fica, S. M., Lin, P.-C., Newman, A. J., & Nagai, K. (2015). CryoEM structures of two spliceosomal complexes: starter and dessert at the spliceosome feast. Current Opinion in Structural Biology, 36, 48–57.
A summary of the first two structures of the spliceosome. The structure was in perfect agreement with previous biochemical studies of the active site.
Book chapters
-
Fica, Sebastian M., Small, E.C., Mefford, M., and Staley, J. P., Mechanistic Insights into Mammalian Pre-mRNA Splicing, Posttranscriptional Gene Regulation: RNA Processing in Eukaryotes, Jane Wu Ed, First Edition, Wiley (2013).